Omics-LHV, West Nile Experiment WCD003 The purpose of this West Nile experiment was to obtain samples for omics analysis in mouse dendritic cell response to wild-type West Nile virus (WNV). Overall Design: Mouse dendritic cells (2 x 10^5) were treated with wild-type WNV and collected in parallel...
Filter results
Category
- Biology (44)
- Scientific Discovery (44)
- Human Health (30)
- Earth System Science (12)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Integrative Omics (3)
- Microbiome Science (2)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- National Security (1)
Content type
Tags
- (-) Viruses (27)
- (-) Microbiome (8)
- (-) Machine Learning (7)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Omics (26)
- Health (23)
- Virus (23)
- Soil Microbiology (22)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (14)
- Synthetic (14)
- sequencing (13)
- West Nile virus (13)
- Genomics (12)
- Ebola (11)
- Influenza A (11)
- Metagenomics (10)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Resource Metadata (9)
- Proteomics (8)
- Microarray (7)
Omics-LHV, West Nile Experiment WCN004 The purpose of this West Nile experiment was to obtain samples for omics analysis in mouse cerebral cortex neurons in response to wild-type West Nile Virus (WNV; WNV-NY99 382) and mutant WNV-E218A (WNV-NY99 382) viral infection. Overall Design: Mouse cortical...
Category
Omics-LHV, West Nile Experiment WGCN004 The purpose of this West Nile experiment was to obtain samples for omics analysis in primary mouse granule neuron cells infected with wild type West Nile virus (WNV-NY99 clone 382, WNVWT) and mutant virus (WNVE218A). Overall Design: Granule cell neurons from...
Category
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
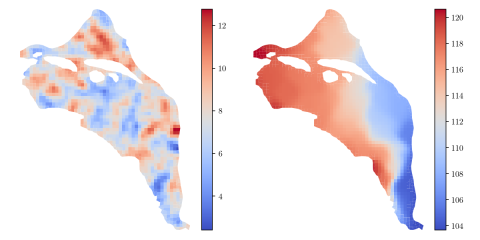
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
Category
"Moisture modulates soil reservoirs of active DNA and RNA viruses" Soil is known to harbor viruses, but the majority are uncharacterized and their responses to environmental changes are unknown. Here, we used a multi-omics approach (metagenomics, metatranscriptomics and metaproteomics) to detect...
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3
PerCon SFA, Co-Investigator Vivian Lin earned her PhD in organic chemistry from the University of California, Berkeley with Professor Chris Chang, developing fluorescent probes for imaging redox active small molecules. Afterward, she traveled to Switzerland for a postdoctoral fellowship in the...
Category
Category
Category
Category
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...