Filter results
Content type
Tags
- (-) Mus musculus (18)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Viruses (26)
- Omics (25)
- Health (23)
- Virus (23)
- Soil Microbiology (21)
- MERS-CoV (18)
- Mass Spectrometry (14)
- Synthetic (14)
- sequencing (13)
- West Nile virus (13)
- Genomics (12)
- Ebola (11)
- Influenza A (11)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Metagenomics (9)
- Resource Metadata (9)
- Microbiome (8)
- Proteomics (8)
- Machine Learning (7)
- Microarray (7)
Category
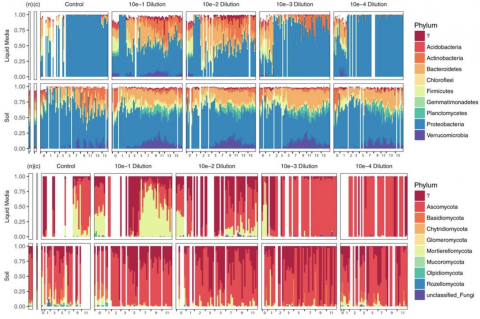
Soil fungi facilitate the translocation of inorganic nutrients from soil minerals to other microorganisms and plants. This ability is particularly advantageous in impoverished soils, because fungal mycelial networks can bridge otherwise spatially disconnected and inaccessible nutrient hotspots...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to MERS-CoV Virus Infection Background Middle East Respiratory Syndrome coronavirus ( MERS-CoV ), part of the Coronaviridae family, is classified as a Category C priority pathogen by the National Institute...
Category
Datasets
15
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Influenza A Virus Infection Background Influenza A virus ( IAV ) is a high risk biological agent belonging to the Orthomyxoviridae family is classified as a Category C priority pathogen by the National...
Category
Datasets
8
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to West Nile Virus Infection Background West Nile virus ( WNV ) belongs to the mosquito-borne Flaviviridae family and is classified as a Category A priority pathogen by the National Institute of Allergy and...
Category
Datasets
11
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MM001 The purpose of this experiment was to evaluate the host response to wild-type MERS-CoV virus infection. Sample data was obtained from primary mouse (strain C57BL/6J) whole lung for mRNA, proteomics, metabolomics, and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment IM102 The purpose of this experiment was to evaluate the mouse host response to Influenza A virus (subtype H7N9) wild-type strain Influenza A/Anhui/1/2013 (AH1-WT) virus and mutants NS1-103F/106M (AH1-F/M) and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment IM103 The purpose of this experiment was to evaluate the host response to Influenza A virus (subtype H5N1) wild-type strain Influenza A/Vietnam/1203/2004 (VN1203) virus, mutant VN1203-NS1trunc124, and mock...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCB001 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCT001 The purpose of this experiment was to evaluate the host responseto West Nile virus (WNV-NY99) wild-type (strain 382) and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain C57BL...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment IM101 The purpose of this experiment was to evaluate the host mouse response to Influenza A virus (subtype H5N1) wild-type strain Influenza A/Vietnam/1203/2004, Influenza A/Vietnam/1203/2004 mutant strains PB2-627E...
Category
Category
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WGCN003 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain...