This data package consists of data from human islets treated for 48 h with cytokines (IFNγ, TNFα, and/or IL1β), thapsigargin (TG), brefeldin A (BFA), or DMSO for the control (CTRL) and submitted for single-cell RNA-seq (n=5) and single-nucleus multi-omics sequencing (snMultiomics-seq, n=1) ( Table 1...
Filter results
Category
- (-) Integrative Omics (96)
- (-) Data Analytics & Machine Learning (9)
- Scientific Discovery (406)
- Biology (287)
- Earth System Science (169)
- Human Health (113)
- Microbiome Science (50)
- National Security (32)
- Computational Research (25)
- Computing & Analytics (18)
- Energy Resiliency (16)
- Chemical & Biological Signatures Science (12)
- Weapons of Mass Effect (12)
- Materials Science (11)
- Chemistry (10)
- Renewable Energy (8)
- Computational Mathematics & Statistics (7)
- Data Analytics & Machine Learning (7)
- Atmospheric Science (6)
- Ecosystem Science (6)
- Visual Analytics (6)
- Coastal Science (4)
- Energy Storage (4)
- Plant Science (4)
- Solar Energy (4)
- Bioenergy Technologies (3)
- Energy Efficiency (3)
- Transportation (3)
- Cybersecurity (2)
- Distribution (2)
- Electric Grid Modernization (2)
- Grid Cybersecurity (2)
- Subsurface Science (2)
- Water Power (2)
- Wind Energy (2)
- Advanced Lighting (1)
- Computational Mathematics & Statistics (1)
- Environmental Management (1)
- Federal Buildings (1)
- Geothermal Energy (1)
- Grid Analytics (1)
- Grid Energy Storage (1)
- High-Performance Computing (1)
- Terrestrial Aquatics (1)
- Vehicle Technologies (1)
- Waste Processing (1)
Tags
- Virology (55)
- Immune Response (51)
- Time Sampled Measurement Datasets (50)
- Differential Expression Analysis (46)
- Gene expression profile data (45)
- Homo sapiens (34)
- Multi-Omics (32)
- Mass spectrometry data (30)
- Mus musculus (19)
- MERS-CoV (18)
- West Nile virus (13)
- Influenza A (11)
- Ebola (9)
- Omics (9)
- Mass Spectrometry (8)
- Resource Metadata (7)
- Type 1 Diabetes (7)
- Human Interferon (6)
- Microarray (6)
- Omics-LHV Project (6)
- Proteomics (6)
- Autoimmunity (5)
- Machine Learning (5)
- Biomarkers (4)
- Mass Spectrometer (4)
- Mass spectrometry-based Omics (4)
- Molecular Profiling (4)
- Imaging (3)
- metabolomics (3)
- Microscopy (3)
This data package consists of data from MIN6 cells treated with IL-1β + IFNγ + TNFα + palmitate for 24 h and submitted to proteomics analysis, and the same treatment for 8 h for metabolomics analysis (n=4) ( Table 1 ). This experimental design was set to study the possible roles of free fatty acids...
Category
This data package consists of data from human islets (n=5) treated with IFNγ + IL-1β with or without difluoromethylornithine (DFMO) and submitted to RNA-seq and proteomics analysis ( Table 1 ). DFMO is an inhibitor of ornithine decarboxylase (ODC), an enzyme of the polyamine pathway. Therefore, this...
Category
This data package consists of data from human islets and EndoC-βH1 cells treated with IFNα for 2, 8, or 24 h and submitted to RNA-seq (human islets: n=6 & EndoC-βH1: n=5) and ATAC-seq (n=4). In parallel, a batch of cells (n=4) had proteomics, lipidomics, and metabolomics performed in the same...
Category
This data package consists of RNA-seq data from human islets (n=10) treated with IFNγ + IL-1β for 24 h ( Table 1 ). Data contributors: Wenting Wu & Carmella Evans-Molina: Center for Diabetes and Metabolic Diseases, Indiana University School of Medicine, Indianapolis, IN, USA. Farooq Syed: Center for...
Category
This data package consists of data from human islets (n=5) treated with IFNγ + IL-1β for 48 h and submitted to RNA-seq analysis ( Table 1 ). Data contributors: Arturo Roca-Rivada, Xiaoyan Yi & Décio L. Eizirik: ULB Center for Diabetes Research, Medical Faculty, Université Libre de Bruxelles...
Category
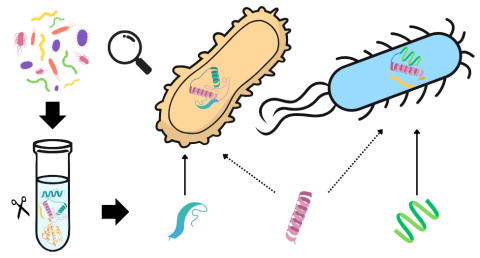
This data package contains processed LC-MS proteomics results and analysis scripts associated with the paper "Assessing Degenerate Peptide Resolution Methods using a Ground Truth Dataset". In this study, we designed an artificial microbial community to create a ground truth, simulated metaproteomic...
Category
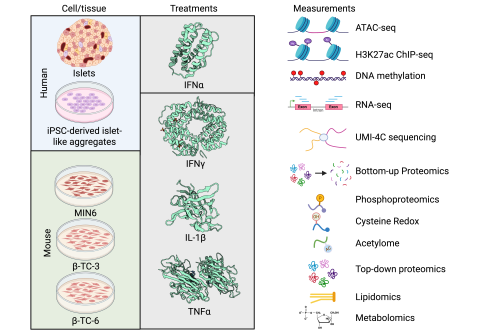
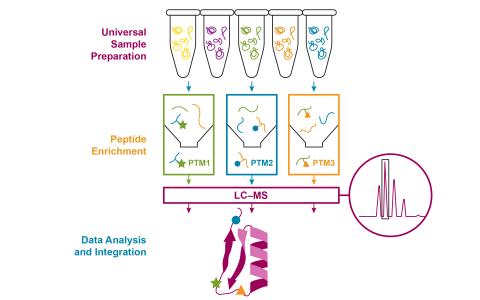
Dataset Description The goal of the experiment was to demonstrate that the optimized multiplexed multi-PTM profiling workflow can comprehensively and quantitatively capture dynamic changes in protein abundance, cysteine oxidation, phosphorylation, and acetylation in cytokine-induced inflammatory...
Category
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...
A total of 172 children from the DAISY study with multiple plasma samples collected over time, with up to 23 years of follow-up, were characterized via proteomics analysis. Of the children there were 40 controls and 132 cases. All 132 cases had measurements across time relative to IA. Sampling was...
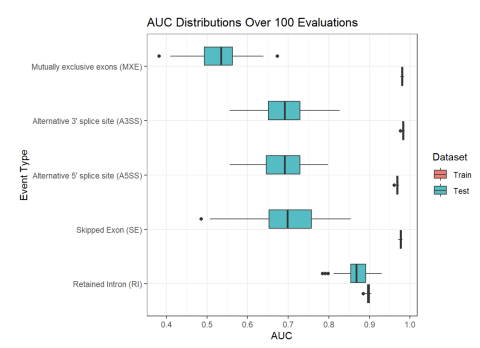
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL005 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to MERS-CoV virus infection. Samples were obtained from human lung adenocarcinoma cells (Calu-3) and processed for...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL004 Metadata The purpose of this experiment was to evaluate the host response to MERS-CoV virus infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and processed for genome binding/occupancy...
Category
Omics-LHV, West Nile Experiment WGCN004 The purpose of this West Nile experiment was to obtain samples for omics analysis in primary mouse granule neuron cells infected with wild type West Nile virus (WNV-NY99 clone 382, WNVWT) and mutant virus (WNVE218A). Overall Design: Granule cell neurons from...
Category
Last updated on 2025-01-16T22:12:33+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...