The data package consists of data from iPSC: HEL115.6-derived islet-like aggregates treated with IFNα for 24 h and submitted to RNA-seq (n=5) and scRNA-seq (n=5). ( Table 1 ). Data contributors: Arturo Roca-Rivada, Xiaoyan Yi & Décio L. Eizirik: ULB Center for Diabetes Research, Medical Faculty...
Filter results
Category
- (-) Integrative Omics (97)
- Scientific Discovery (407)
- Biology (288)
- Earth System Science (169)
- Human Health (113)
- Microbiome Science (50)
- National Security (32)
- Computational Research (25)
- Energy Resiliency (22)
- Computing & Analytics (18)
- Renewable Energy (14)
- Chemical & Biological Signatures Science (12)
- Weapons of Mass Effect (12)
- Materials Science (11)
- Chemistry (10)
- Data Analytics & Machine Learning (9)
- Water Power (8)
- Computational Mathematics & Statistics (7)
- Data Analytics & Machine Learning (7)
- Atmospheric Science (6)
- Ecosystem Science (6)
- Visual Analytics (6)
- Coastal Science (4)
- Energy Storage (4)
- Plant Science (4)
- Solar Energy (4)
- Bioenergy Technologies (3)
- Energy Efficiency (3)
- Transportation (3)
- Cybersecurity (2)
- Distribution (2)
- Electric Grid Modernization (2)
- Grid Cybersecurity (2)
- Subsurface Science (2)
- Wind Energy (2)
- Advanced Lighting (1)
- Computational Mathematics & Statistics (1)
- Environmental Management (1)
- Federal Buildings (1)
- Geothermal Energy (1)
- Grid Analytics (1)
- Grid Energy Storage (1)
- High-Performance Computing (1)
- Terrestrial Aquatics (1)
- Vehicle Technologies (1)
- Waste Processing (1)
Tags
- Virology (55)
- Immune Response (51)
- Time Sampled Measurement Datasets (50)
- Differential Expression Analysis (46)
- Gene expression profile data (45)
- Homo sapiens (34)
- Multi-Omics (32)
- Mass spectrometry data (30)
- Mus musculus (19)
- MERS-CoV (18)
- West Nile virus (13)
- Influenza A (11)
- Ebola (9)
- Omics (9)
- Mass Spectrometry (8)
- Resource Metadata (7)
- Human Interferon (6)
- Microarray (6)
- Omics-LHV Project (6)
- Proteomics (6)
- Mass Spectrometer (4)
- Imaging (3)
- metabolomics (3)
- Microscopy (3)
- Response to type I interferon (3)
- Epigenetics (2)
- Lethal Human Viruses (2)
- Lipidomics (2)
- Processed Data (2)
- Viral Experiment (2)
This data package consists of data from human islets (n=8) treated with IFNγ + IL-1β for 24 h and submitted to phosphoproteomics ( Table 1 ). Data contributor: Wei-Jun Qian: Biological Sciences Division, Pacific Northwest National Laboratory, Richland, WA, 99354, USA. Data repository: PXD011857...
Category
The data package consists of RNA-seq datasets of EndoC-βH1 cells treated in parallel with IFNα, IFNγ, IL-1β and TNFα for 24 h and submitted to RNA-seq (n=4) ( Table 1 ) Data contributors: Arturo Roca-Rivada, Xiaoyan Yi & Décio L. Eizirik: ULB Center for Diabetes Research, Medical Faculty, Université...
Category
It consists of human islets (n=10, subdivided into 2 batches) treated with IFNγ + IL-1β for 24 h and submitted to proteomics, lipidomics and metabolomics in the same samples ( Table 1 ) Data contributors: Ernesto S. Nakayasu, Jennifer Kyle, Young-Mo Kim, Soumyadeep Sarkar & Thomas O. Metz...
Category
This data package consists of data from human islets (n=6) treated with IFNγ + IL-1β for 24 h and submitted to top-down proteomics ( Table 1 ). Data contributors: Ashley Ives, Wei-Jun Qian & James Fulcher: Biological Sciences Division, Pacific Northwest National Laboratory, Richland, WA, 99354, USA...
Category
This data package consists of data from MIN6 cells (n=4) treated with IL-1β + IFNγ + TNFα for 24 h post RNAi targeting 85/88 kDa calcium-independent phospholipase A2 ( Pla2g6 ) and submitted to proteomics analysis ( Table 1 ). Pla2g6 is an enzyme that metabolizes phosphatidylcholine (PCs) into...
Category
This data package consists of data from MIN6 cells (n=4) treated with IL-1β + IFNγ + TNFα and RNAi targeting Poly(ADP)ribosyl polymerase 12 ( Parp12 ) for 24 h and submitted to proteomic analyses ( Table 1 ). PARP12 is an interferon-induced protein known to target viral RNA to stress granules. It...
Category
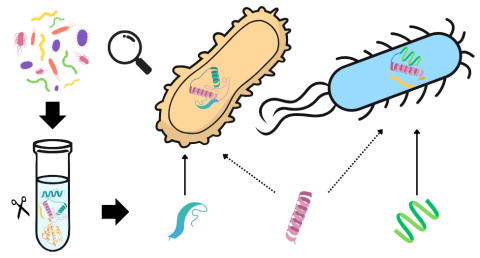
This data package contains processed LC-MS proteomics results and analysis scripts associated with the paper "Assessing Degenerate Peptide Resolution Methods using a Ground Truth Dataset". In this study, we designed an artificial microbial community to create a ground truth, simulated metaproteomic...
Category
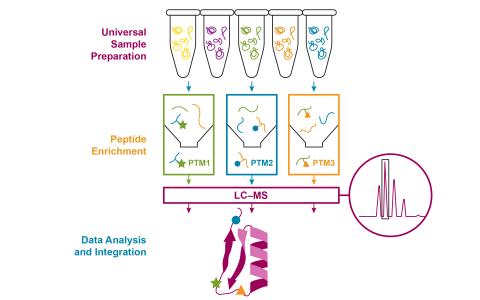
Dataset Description The goal of the experiment was to demonstrate that the optimized multiplexed multi-PTM profiling workflow can comprehensively and quantitatively capture dynamic changes in protein abundance, cysteine oxidation, phosphorylation, and acetylation in cytokine-induced inflammatory...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL005 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to MERS-CoV virus infection. Samples were obtained from human lung adenocarcinoma cells (Calu-3) and processed for...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL004 Metadata The purpose of this experiment was to evaluate the host response to MERS-CoV virus infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and processed for genome binding/occupancy...
Category
Omics-LHV, West Nile Experiment WGCN004 The purpose of this West Nile experiment was to obtain samples for omics analysis in primary mouse granule neuron cells infected with wild type West Nile virus (WNV-NY99 clone 382, WNVWT) and mutant virus (WNVE218A). Overall Design: Granule cell neurons from...
Category
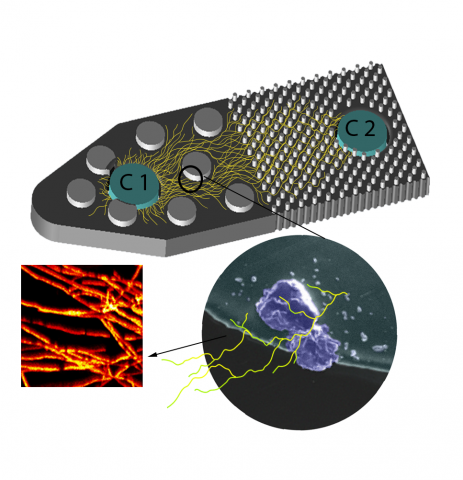
Last updated on 2025-01-16T22:12:33+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL105 Metadata The purpose of this experiment was to evaluate the host epigenetic response to Influenza A virus (subtype H5N1) infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL106 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to Influenza A virus (subtype H5N1) infection. Samples were obtained from human lung adenocarcinoma cell line (Calu...