Description
Dr. Nelson is a computational scientist at PNNL. Since coming to PNNL, he has focused on methods to determine precise understanding of microbial community structure and function through the resolution of individual genome sequences from metagenomic data, and exploration of variation between individuals in microbial populations. Recently, he has focused on integrating proteomics and metabolomics with genomic analysis to achieve a systems biology modeling approach to examining microbial ecology in complex communities such as soil and human gut microbiome.
Expertise
Projects (2)
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
5
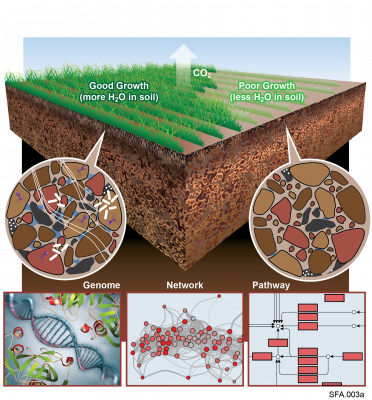
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
45