Filter results
Category
- (-) Earth System Science (11)
- (-) Microbiome Science (3)
- Scientific Discovery (18)
- Biology (17)
- Computational Research (6)
- Data Analytics & Machine Learning (5)
- Human Health (5)
- Computational Mathematics & Statistics (2)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- National Security (1)
Content type
Tags
- (-) Metagenomics (9)
- (-) Output Databases (2)
- Omics (22)
- Soil Microbiology (21)
- sequencing (13)
- Genomics (12)
- High Throughput Sequencing (9)
- PerCon SFA (9)
- Microbiome (8)
- Fungi (6)
- Imaging (6)
- Mass Spectrometry (6)
- Mass Spectrometer (5)
- Sequencer System (5)
- Synthetic Biology (5)
- metagenomics (4)
- Microscopy (4)
- Proteomics (4)
- Sequencing (4)
- soil microbiology (4)
- Spectroscopy (4)
- Amplicon Sequencing (3)
- Climate Change (3)
- IAREC (3)
- Mass spectrometry data (3)
- metabolomics (3)
- Viruses (3)
- Whole Genome Sequencing (3)
- microbiome stability (2)
- species volatility (2)
"DNA Viral Diversity, Abundance, and Functional Potential Vary across Grassland Soils with a Range of Historical Moisture Regimes" Soil viruses are abundant, but the influence of the environment and climate on soil viruses remains poorly understood. Here, we addressed this gap by comparing the...
Category
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
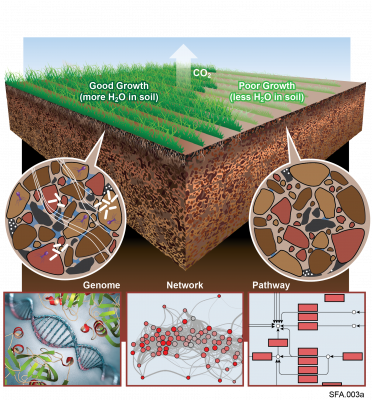
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Category
Category
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. WA-TmG.2.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG2/1770324 Soil samples were collected in triplicate in the fall of 2017 across the three grassland...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. KS-TmG.2.0 (Metagenome, KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/KSTmG2/1770332 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. Iso-VIG14.1.0 (Metagenome Derived Viral Genomes, WA/IA/KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/IsoVIG14/1770369 Soil samples were collected in triplicate in the Fall of 2017...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. IA-TmG.2.0 (Metagenome, IA). [Data Set] PNNL DataHub. https://doi.org/10.25584/IATmG2/1770333 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
These GCAM v4.3 SSP-RCP-GCM Output Databases are made available under the Open Data Commons Attribution License: http://opendatacommons.org/licenses/by/1.0/ . GCAM v4.3 SSP-RCP-GCM plausible solution databases. Supplemental dataset to: Graham N.T., M.I. Hejazi, M. Chen, E. Davies, J.A. Edmonds, S.H...