Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...
Filter results
Category
- (-) Scientific Discovery (25)
- Biology (25)
- Human Health (20)
- Integrative Omics (15)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Earth System Science (3)
- Microbiome Science (3)
- National Security (3)
- Chemical & Biological Signatures Science (2)
- Weapons of Mass Effect (2)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
Content type
Tags
- (-) Ebola (9)
- (-) Proteomics (8)
- Virology (77)
- Immune Response (51)
- Time Sampled Measurement Datasets (50)
- Differential Expression Analysis (46)
- Gene expression profile data (45)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Viruses (27)
- Omics (26)
- Health (23)
- Virus (23)
- Soil Microbiology (22)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (14)
- sequencing (13)
- West Nile virus (13)
- Genomics (12)
- Influenza A (11)
- Metagenomics (10)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Microbiome (8)
- Microarray (7)
- Resource Metadata (7)
- Synthetic Biology (7)
- Imaging (6)
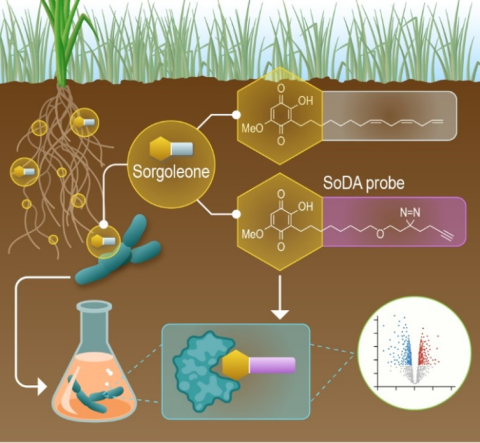
Last updated on 2024-06-04T17:11:36+00:00 by LN Anderson PerCon SFA: Profiling sorghum-microbe interactions with a specialized photoaffinity probe identifies key sorgoleone binders in Acinetobacter pitti Mass spectrometry-based global proteome analysis and SoDA-PAL photoaffinity probe labeled...
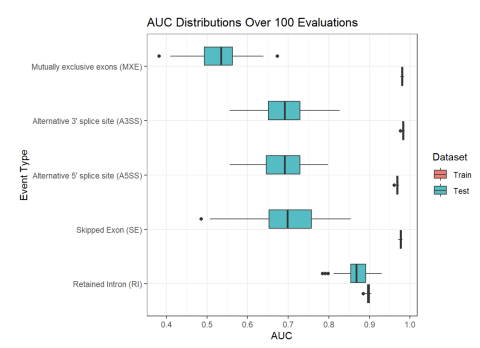
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
Biomedical Resilience & Readiness in Adverse Operating Environments (BRAVE) Project: Exhaled Breath Condensate (EBC) TMT Proteomic Transformation Data Exhaled breath condensate (EBC) represents a low-cost and non-invasive means of examining respiratory health. EBC has been used to discover and...
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EH001 The purpose of this experiment was to evaluate the human patient peripheral blood mononuclear cells (PBMC) response to Zaire Ebola virus (strain Makona) infection during the 2013-2016 epidemic in West Africa...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EU937001 The purpose of this experiment was to evaluate the human host response to wild-type Zaire Ebola virus (strain Mayinga) and mutant virus infection. Samples were obtained from human histiocytic lymphoma cells...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EHUVEC001 The purpose of this experiment was to evaluate the human host response to wild-type Zaire Ebola virus (strain Mayinga) and mutant virus infection in VP30 expression background. Sample data was obtained from...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EHUH003 The purpose of this experiment was to evaluate the human host response to wild-type Zaire Ebola virus (strain Mayinga) infection. Samples were obtained from human hepatoma carcinoma cells (HUH-7) infected with...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EHUH002 The purpose of this experiment was to evaluate the human host response to wild-type Zaire Ebola virus (strain Mayinga) and mutant virus infection. Samples were obtained from human hepatoma carcinoma cells (HUH-7)...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EHUH001 The purpose of this experiment was to evaluate the human host response to wild-type Zaire Ebola virus (strain Mayinga) and mutant virus infection. Samples were obtained from human hepatoma carcinoma cells (HUH-7)...
Category
MERS-CoV Experiment MDC001 Processed Omics Data Unavailable This experiment evaluated primary human dendritic cells infected with a wild type MERS-CoV (icMERS) virus. Related Experimental Data BioProject: PRJNA315103 GEO: GSE79172 (mRNA transcriptome response) Acknowledgment of Federal Funding The...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Interferon Experiment IFNaHUH001 The purpose of this experiment was to evaluate the human host cellular response to treatment with and without interferon alpha/beta (IFNα/β) treatment. Sample time course data was obtained from human hepatoma...
Category
Human infections caused by viral pathogens trigger a complex gamut of host responses that limit disease, resolve infection, generate immunity, and contribute to severe disease or death. Here, we present experimental methods and multi-omics data capture approaches representing the global host...
Category
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1