Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCN002 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain C57BL...
Filter results
Content type
Dataset Type
- (-) Transcriptomics (51)
- (-) Amplicon (16S, ITS) (15)
- (-) Staff (2)
- Omics (86)
- Proteomics (39)
- Sequencing (39)
- Metabolomics (28)
- Lipidomics (26)
- Simulation Results (17)
- Metagenomics (9)
- Microarray (9)
- Computed Analysis (4)
- Genomics (4)
- ChIP-seq (2)
- Geospacial (2)
- Cybersecurity (1)
- Electromagnetic Spectrum (1)
- Mass Spectra (1)
- Microscopy (1)
- Phenomics (1)
- Whole genome sequencing (WGS) (1)
Tags
- Virology (49)
- Gene expression profile data (40)
- Immune Response (40)
- Time Sampled Measurement Datasets (37)
- Differential Expression Analysis (35)
- Homo sapiens (24)
- Mass spectrometry data (18)
- Multi-Omics (17)
- MERS-CoV (13)
- Mus musculus (13)
- West Nile virus (9)
- Health (8)
- Virus (8)
- Viruses (8)
- Ebola (7)
- Influenza A (7)
- Microarray (5)
- Resource Metadata (5)
- Human Interferon (4)
- Response to type I interferon (3)
- Soil Microbiology (3)
- Amplicon Sequencing (2)
- Response to type I & type II interferon (2)
- sequencing (2)
- IAREC (1)
- Mass Spectrometry (1)
- Microbiome (1)
- microbiome stability (1)
- Omics (1)
- species volatility (1)
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCN003 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain C57BL...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WLN002 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain...
Category
Omics Lethal Human Virus, SARS-CoV Experiment SM001 New uploads pending The purpose of this experiment was to evaluate the human host response to Severe Acute Respiratory Syndrome coronavirus (SARS-CoV) wild-type virus. Sample data was obtained for 20 week-old C57BL/6J mouse lung tissue infected...
Category
The VAST 2009 Challenge scenario concerned a fictitious, cyber security event. An employee leaked important information from an embassy to a criminal organization. Participants were asked to discover the identity ofthe employee and the structure of the criminal organization. Participants were...
Category
This data set provides the ingrowth peat microbial biomass carbon (MBC) and nitrogen (MBN), extractable organic carbon (EOC) and extractable nitrogen (EN) for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatland Responses Under Changing Environments...
Category
This data set provides the ingrowth peat extracellular enzyme potential (EE) for before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). EE potential was quantified and calculated following a...
Category
This data set provides the 16S microbial community composition via DNA and cDNA sequence analyses at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were extracted using a...
Category
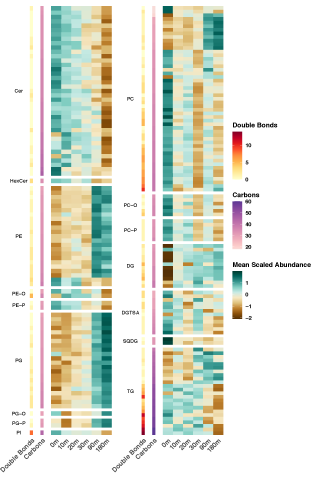
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
This data set provides ITS fungal community composition via DNA sequence analysis at the time of peat coring at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the Spruce and...
Category
This data set provides the 16S microbial community composition via DNA sequence analysis from ingrowth peat and sand cores at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the...
Category
This data set provides the ITS fungal community composition via DNA sequence analysis from sand and peat ingrowth cores at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the Spruce...
Category
This data set provides the peat microbial biomass carbon (MBC) and nitrogen (MBN), extractable organic carbon (EOC) and extractable nitrogen (EN) at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatland Responses Under...
Category
This data set provides the 16S microbial community composition of peat and sand ingrowth cores via DNA and cDNA sequence analysis before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were...
Category
This data set provides ITS fungal community composition via DNA and cDNA sequence analysis at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were extracted using a Qiagen...