Filter results
Content type
Tags
- (-) Genomics (7)
- Virology (77)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (29)
- Multi-Omics (28)
- Viruses (24)
- Health (21)
- Soil Microbiology (21)
- Virus (21)
- MERS-CoV (18)
- Mus musculus (18)
- Synthetic (14)
- sequencing (13)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- Omics (10)
- Metagenomics (9)
- Microbiome (8)
- PerCon SFA (8)
- Resource Metadata (8)
- Machine Learning (7)
- Fungi (6)
- Human Interferon (6)
- Mass Spectrometry (6)
- Microarray (6)
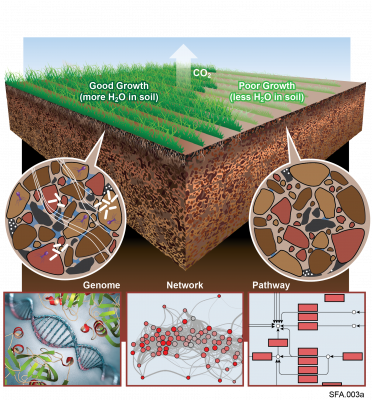
Viral communities detected from three large grassland soil metagenomes with historically different precipitation moisture regimes.
Category
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Understanding molecular-scale factors governing the precipitation of aluminum hydroxides, such as gibbsite, under alkaline conditions is important for the formation of laterite deposits, as well as aluminum processing. However, mechanisms enabling tetrahedral aluminate ions to assemble into...
Electrolyte solutions in alkaline nuclear waste contain aluminate, hydroxide, nitrate and nitrite with sodium as the predominant counterion. The salts of these ions are highly soluble, so the liquids are highly concentrated. This study found that there is a substantial incompatibility between the...
The recently developed real-time nuclear–electronic orbital (RT-NEO) approach provides an elegant framework for treating electrons and selected nuclei, typically protons, quantum mechanically in nonequilibrium dynamical processes. However, the RT-NEO approach neglects the motion of the other nuclei...
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. Iso-VIG14.1.0 (Metagenome Derived Viral Genomes, WA/IA/KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/IsoVIG14/1770369 Soil samples were collected in triplicate in the Fall of 2017...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes. [Data Set] PNNL DataHub. https://doi.org/10.25584/PNNLDH/1986536 Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes...
Category
Last updated on 2023-05-31T16:35:53+00:00 by LN Anderson PerCon SFA: Sequencing of Sorgoleone Promoting Rhizobacteria Isolates Whole genome sequencing (WGS) of sorgoleone utilizing rhizobacteria strains Pseudomonas sorgoleonovorans SO81 , Burkholderia anthina SO82 , and Acinetobacter pittii SO1 , as...