Systems Virology Lethal Human Virus, SARS-CoV Experiment SM012 New uploads pending The purpose of this SARS experiment was to obtain samples for transcriptome and proteome analysis in C57Bl/6 mice infected with SARS MA15 or SARS deltaORF6 mutant viruses. Overall Design: Twenty-week-old C57BL/6 mice...
Filter results
Category
- Biology (40)
- Scientific Discovery (40)
- Human Health (38)
- Integrative Omics (29)
- Computational Mathematics & Statistics (2)
- Computational Research (2)
- Data Analytics & Machine Learning (2)
- Chemical & Biological Signatures Science (1)
- Earth System Science (1)
- Microbiome Science (1)
- National Security (1)
- Weapons of Mass Effect (1)
Content type
Dataset Type
- (-) Proteomics (39)
- (-) ChIP-seq (2)
- Omics (86)
- Transcriptomics (51)
- Sequencing (39)
- Metabolomics (28)
- Lipidomics (26)
- Simulation Results (17)
- Amplicon (16S, ITS) (15)
- Metagenomics (9)
- Microarray (9)
- Genomics (4)
- Computed Analysis (3)
- Geospacial (2)
- Staff (2)
- Cybersecurity (1)
- Electromagnetic Spectrum (1)
- Mass Spectra (1)
- Microscopy (1)
- Phenomics (1)
- Whole genome sequencing (WGS) (1)
Tags
- Virology (35)
- Immune Response (26)
- Time Sampled Measurement Datasets (26)
- Mass spectrometry data (25)
- Differential Expression Analysis (24)
- Multi-Omics (23)
- Gene expression profile data (20)
- Homo sapiens (18)
- MERS-CoV (12)
- Health (8)
- Mus musculus (8)
- Virus (8)
- Viruses (8)
- Influenza A (7)
- West Nile virus (5)
- Ebola (2)
- Epigenetics (2)
- Proteomics (2)
- Resource Metadata (2)
- EBC (1)
- Fungi (1)
- Machine Learning (1)
- Mass Spectrometry (1)
- Microscopy (1)
- mineral weathering (1)
- Omics (1)
- Quantification (1)
- Soil Microbiology (1)
- Spectroscopy (1)
- TMT (1)
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EH001 The purpose of this experiment was to evaluate the human patient peripheral blood mononuclear cells (PBMC) response to Zaire Ebola virus (strain Makona) infection during the 2013-2016 epidemic in West Africa...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL003 The purpose of this experiment was to evaluate the human host response to wild-type infectious clone of Middle Eastern Respiratory Syndrome coronavirus (icMERS-CoV) infection and mockulum. Sample data was obtained...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
Biomedical Resilience & Readiness in Adverse Operating Environments (BRAVE) Project: Exhaled Breath Condensate (EBC) TMT Proteomic Transformation Data Exhaled breath condensate (EBC) represents a low-cost and non-invasive means of examining respiratory health. EBC has been used to discover and...
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL105 Metadata The purpose of this experiment was to evaluate the host epigenetic response to Influenza A virus (subtype H5N1) infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL106 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to Influenza A virus (subtype H5N1) infection. Samples were obtained from human lung adenocarcinoma cell line (Calu...
Category
A pre-trained convolutional neural network was fine-tuned for three separate classification tasks, distinguishing 2D images of: 1) single amino acids, 2) protein structural ball and stick images of metalloproteins, and 3) protein structural ball and stick images of metalloenzymes with the metal...
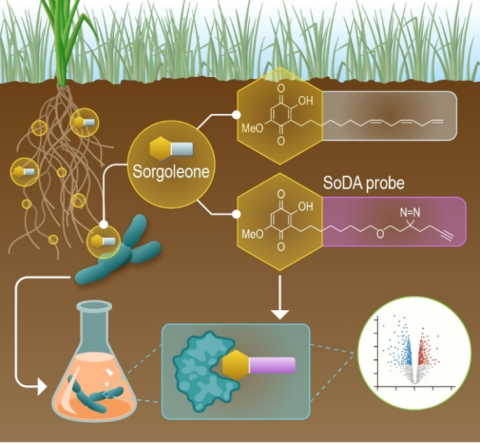
Last updated on 2024-04-19T19:12:08+00:00 by LN Anderson PerCon SFA: Profiling sorghum-microbe interactions with a specialized photoaffinity probe identifies key sorgoleone binders in Acinetobacter pitti Mass spectrometry-based global proteome analysis and SoDA-PAL photoaffinity probe labeled...
A total of 172 children from the DAISY study with multiple plasma samples collected over time, with up to 23 years of follow-up, were characterized via proteomics analysis. Of the children there were 40 controls and 132 cases. All 132 cases had measurements across time relative to IA. Sampling was...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...