Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH001 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to wild-type Ebola viruses Zaire Ebola (ZEBOV '76) and Reston Ebola (REBOV '08) viral infection. Processed Transcriptome...
Filter results
Category
- (-) Biology (67)
- Scientific Discovery (68)
- Integrative Omics (46)
- Human Health (41)
- Earth System Science (26)
- Microbiome Science (10)
- Chemical & Biological Signatures Science (1)
- Chemistry (1)
- Computational Research (1)
- Ecosystem Science (1)
- Materials Science (1)
- National Security (1)
- Plant Science (1)
- Weapons of Mass Effect (1)
Tags
- (-) Multi-Omics (30)
- (-) Omics (24)
- (-) Genomics (12)
- (-) Ebola (11)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Viruses (26)
- Health (23)
- Virus (23)
- Soil Microbiology (21)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (13)
- sequencing (13)
- West Nile virus (13)
- Influenza A (11)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Metagenomics (9)
- Resource Metadata (9)
- Microbiome (8)
- Proteomics (8)
- Microarray (7)
- Synthetic Biology (7)
- Imaging (6)
Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH002 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to response to genetically-reconstructed Zaire Ebola virus infection. Processed Transcriptome Data Unavailable
Category
Category
Viral communities detected from three large grassland soil metagenomes with historically different precipitation moisture regimes.
Category
Soil fungi facilitate the translocation of inorganic nutrients from soil minerals to other microorganisms and plants. This ability is particularly advantageous in impoverished soils, because fungal mycelial networks can bridge otherwise spatially disconnected and inaccessible nutrient hotspots...
Category
"Deconstructing the Soil Microbiome into Reduced-Complexity Functional Modules" The soil microbiome represents one of the most complex microbial communities on the planet, encompassing thousands of taxa and metabolic pathways, rendering holistic analyses computationally intensive and difficult. Here...
Category
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
The Sequel II System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by PacBio , and is designed for high throughput, production-scale sequencing laboratories. Originally released in 2015, the Sequel system provides Single Molecule, Real-Time (SMRT) sequencing core...
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WLN003 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. WA-TmG.2.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG2/1770324 Soil samples were collected in triplicate in the fall of 2017 across the three grassland...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
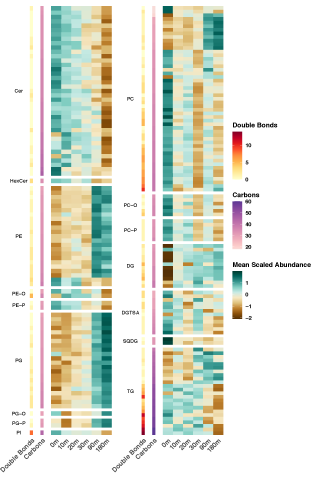
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. Iso-VIG14.1.0 (Metagenome Derived Viral Genomes, WA/IA/KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/IsoVIG14/1770369 Soil samples were collected in triplicate in the Fall of 2017...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. KS-TmG.2.0 (Metagenome, KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/KSTmG2/1770332 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes. [Data Set] PNNL DataHub. https://doi.org/10.25584/PNNLDH/1986536 Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes...