Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH001 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to wild-type Ebola viruses Zaire Ebola (ZEBOV '76) and Reston Ebola (REBOV '08) viral infection. Processed Transcriptome...
Filter results
Category
- Biology (18)
- Scientific Discovery (18)
- Human Health (13)
- Integrative Omics (9)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Earth System Science (5)
- Microbiome Science (2)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- National Security (1)
Tags
- (-) Resource Metadata (8)
- (-) Machine Learning (6)
- (-) Mass Spectrometry (4)
- Virology (69)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (29)
- Multi-Omics (28)
- MERS-CoV (18)
- Mus musculus (18)
- Synthetic (14)
- Viruses (14)
- Health (13)
- Soil Microbiology (13)
- Virus (13)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- sequencing (9)
- Metagenomics (7)
- Omics (7)
- Human Interferon (6)
- Omics-LHV Project (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Genomics (5)
- Microarray (5)
Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH002 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to response to genetically-reconstructed Zaire Ebola virus infection. Processed Transcriptome Data Unavailable
Category
MERS-CoV Experiment MDC001 Processed Omics Data Unavailable This experiment evaluated primary human dendritic cells infected with a wild type MERS-CoV (icMERS) virus. Related Experimental Data BioProject: PRJNA315103 GEO: GSE79172 (mRNA transcriptome response) Acknowledgment of Federal Funding The...
Category
Influenza A Experiment IM104 Processed Omics Data Unavailable This Influenza experiment evaluated mouse lung expression after etoposide treatment and infection with a pandemic H1N1 influenza strain. Related Experimental Data BioProject: PRJNA382278 GEO: GSE97555 (mRNA transcriptome response)...
Category
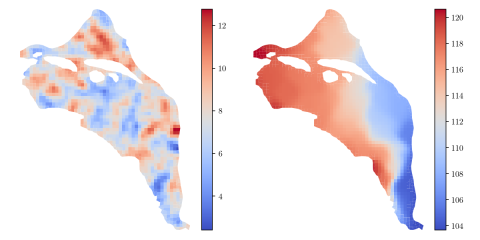
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
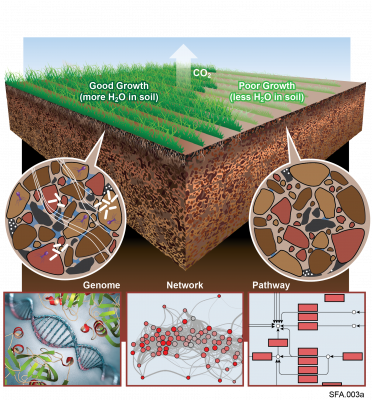
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
3
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3
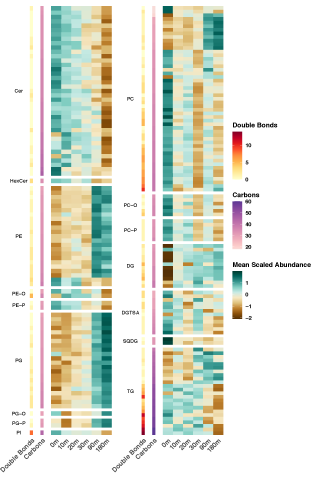
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL105 Metadata The purpose of this experiment was to evaluate the host epigenetic response to Influenza A virus (subtype H5N1) infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes. [Data Set] PNNL DataHub. https://doi.org/10.25584/PNNLDH/1986536 Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL106 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to Influenza A virus (subtype H5N1) infection. Samples were obtained from human lung adenocarcinoma cell line (Calu...