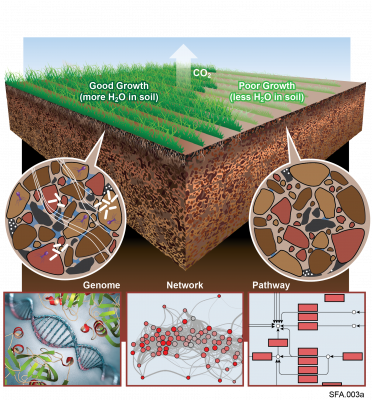
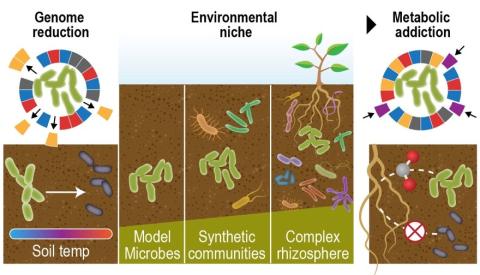
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Category
- (-) Earth System Science (11)
- (-) Computational Research (6)
- Scientific Discovery (25)
- Biology (24)
- Human Health (8)
- Integrative Omics (8)
- Data Analytics & Machine Learning (5)
- Microbiome Science (5)
- Chemistry (3)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Ecosystem Science (1)
- National Security (1)
- Plant Science (1)
Content type
Tags
- (-) Mass Spectrometry (7)
- (-) Machine Learning (5)
- (-) IAREC (3)
- Omics (22)
- Soil Microbiology (21)
- sequencing (13)
- Genomics (10)
- Metagenomics (9)
- Microbiome (8)
- Fungi (6)
- High Throughput Sequencing (6)
- Imaging (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Mass Spectrometer (5)
- Biomarkers (4)
- metagenomics (4)
- Microscopy (4)
- Molecular Profiling (4)
- Sequencer System (4)
- Sequencing (4)
- Software Data Analysis (4)
- soil microbiology (4)
- Spectroscopy (4)
- Climate Change (3)
- metabolomics (3)
- PerCon SFA (3)
- Proteomics (3)
- Synthetic Biology (3)
- Viruses (3)
Pending Review Microbiomes contribute to multiple ecosystem services by transforming organic matter in soil. Extreme shifts in the environment, such as drying-rewetting cycles during drought, can impact microbial metabolism of organic matter by altering their physiology and function. These...
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
pmartR Software Overview The pmartR package provides a single software tool for QC (filtering and normalization), exploratory data analysis (EDA), and statistical analysis (robust to missing data) and includes numerous visualization capabilities of mass spectrometry (MS) omics data (proteomic...
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
3
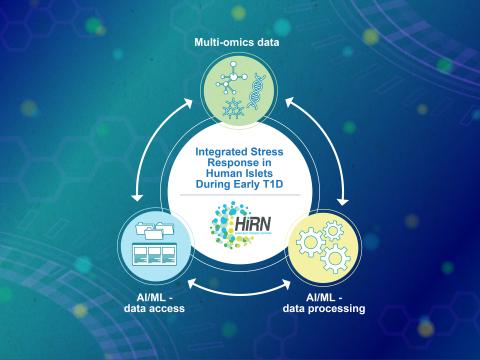
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1
Last updated on 2024-03-03T02:26:52+00:00 by LN Anderson The Thermo Scientific™ Velos Pro™ Orbitrap Mass Spectrometer instrument data source combines advanced mass accuracy with an ultra-high resolution Orbitrap mass analyzer for increased sensitivity, enabling molecular weight determination for...
The IONTOF TOF.SIMS 5 data source is a time-of-flight secondary ion mass spectrometer and powerful surface analysis tool used to investigate scientific questions in biological, environmental, and energy research. Among the most sensitive of surface analysis tools, it uses a high-vacuum technique...
Category
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. WA-TmG.2.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG2/1770324 Soil samples were collected in triplicate in the fall of 2017 across the three grassland...
Category
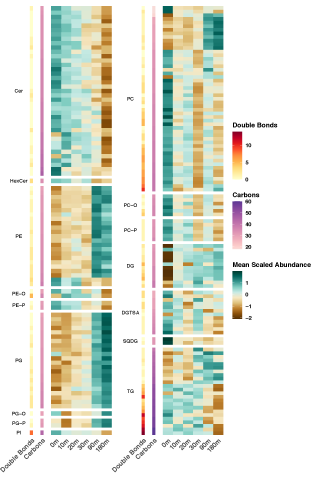
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...