The Illumina MiSeq System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by Illumina , and is designed for sequencing data acquisition using synthesis technology to provide an end-to-end solution (cluster generation, amplification, sequencing, and data analysis) in a...
Filter results
Content type
Tags
- (-) Soil Microbiology (21)
- (-) Genomics (10)
- (-) Synthetic (5)
- Omics (22)
- sequencing (13)
- Metagenomics (9)
- Microbiome (8)
- Fungi (6)
- High Throughput Sequencing (6)
- Imaging (6)
- Mass Spectrometry (6)
- Mass Spectrometer (5)
- Proteomics (5)
- metagenomics (4)
- Microscopy (4)
- Sequencer System (4)
- Sequencing (4)
- soil microbiology (4)
- Spectroscopy (4)
- Climate Change (3)
- IAREC (3)
- metabolomics (3)
- PerCon SFA (3)
- Synthetic Biology (3)
- Viruses (3)
- Cybersecurity (2)
- Lipidomics (2)
- microbiome stability (2)
- omics (2)
- species volatility (2)
The Illumina HiSeq X System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by Illumina , and is designed for high throughput, production-scale sequencing laboratories. Built off the HiSeq 2500 System, harnessing the patterned flow cell technology originally developed...
The Illumina HiSeq 4000 System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by Illumina , and is designed for high throughput, production-scale sequencing laboratories. Built off the HiSeq 2500 System and harnessing the patterned flow cell technology originally...
This data was generated by the organization IvySys. Activities can be phone calls, transactions, or any other type of communications. Most of the files are of the type .edges, .rdf, or .csv; but all can be opened in a text editor. A good introduction to this data can be found in \Tutorial1\MAA...
Category
Category
Category
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Category
Category
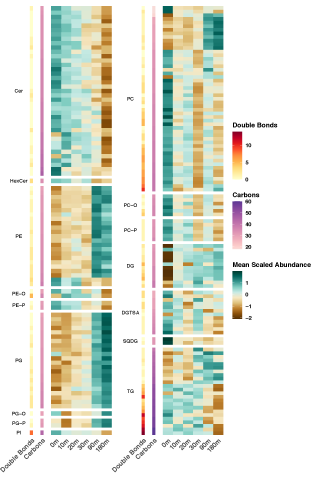
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. WA-TmG.2.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG2/1770324 Soil samples were collected in triplicate in the fall of 2017 across the three grassland...
Category
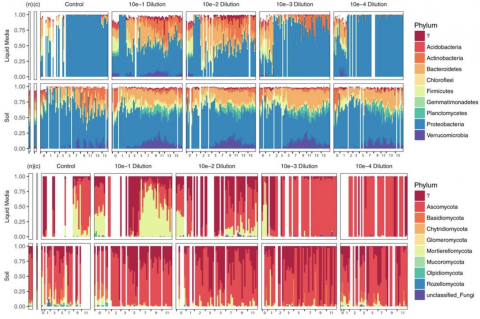
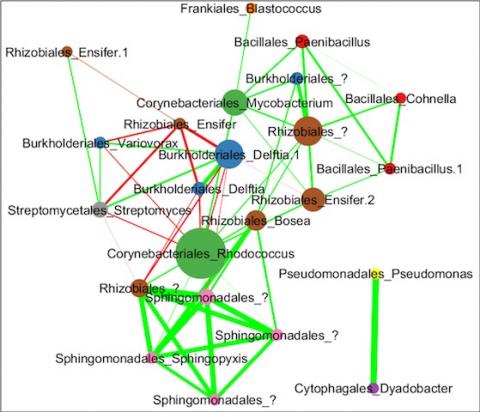
Please cite as : Anderson L.N., J.E. McDermott, and R.S. McClure. 2020. WA-IsoC_MSC1.1.0 (Amplicon 16S rRNA, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WAIsoCMSC1/1635272 The soil microbiome is central to the cycling of carbon and other nutrients and to the promotion of plant growth...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. Iso-VIG14.1.0 (Metagenome Derived Viral Genomes, WA/IA/KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/IsoVIG14/1770369 Soil samples were collected in triplicate in the Fall of 2017...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. KS-TmG.2.0 (Metagenome, KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/KSTmG2/1770332 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...