The innovative Elstar™ electron column forms the basis of the Helios NanoLab’s outstanding high resolution imaging performance. The Elstar features unique technologies, such as constant power lenses for higher thermal stability, electrostatic scanning for faster, higher accurate imaging, and unique...
Filter results
Content type
Tags
- (-) sequencing (13)
- (-) West Nile virus (13)
- (-) Genomics (12)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Viruses (27)
- Omics (26)
- Health (23)
- Virus (23)
- Soil Microbiology (22)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (14)
- Synthetic (14)
- Ebola (11)
- Influenza A (11)
- Metagenomics (10)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Resource Metadata (9)
- Microbiome (8)
- Proteomics (8)
- Machine Learning (7)
- Microarray (7)
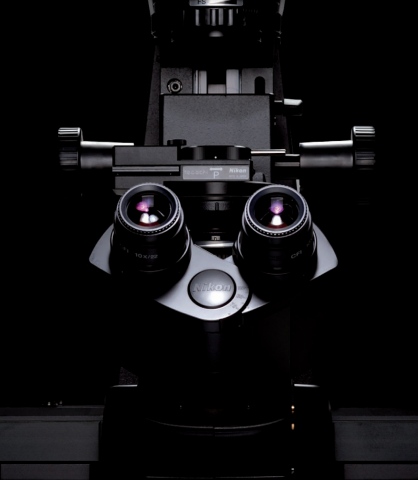
The Nikon Eclipse Ti2-E is a motorized and intelligent model for advanced imaging applications. Compatible with PFS, auto correction collar, and external phase contrast system. The base of choice for live-cell imaging, high-content applications, confocal and super-resolution.As research trends...
Category
Stanford Synchrotron Radiation Lightsource Experimental Station 14-3b is a bending magnet side station dedicated to X-Ray Imaging and Micro X-Ray Absorption Spectroscopy of biological, biomedical, materials, and geological samples. Station 14-3b is equipped with specialized instrumentation for XRF...
Category
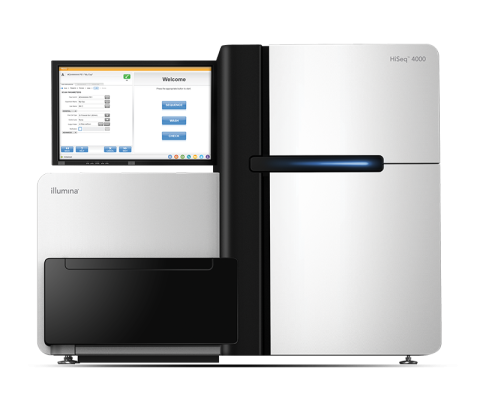
The Illumina HiSeq 4000 System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by Illumina , and is designed for high throughput, production-scale sequencing laboratories. Built off the HiSeq 2500 System and harnessing the patterned flow cell technology originally...
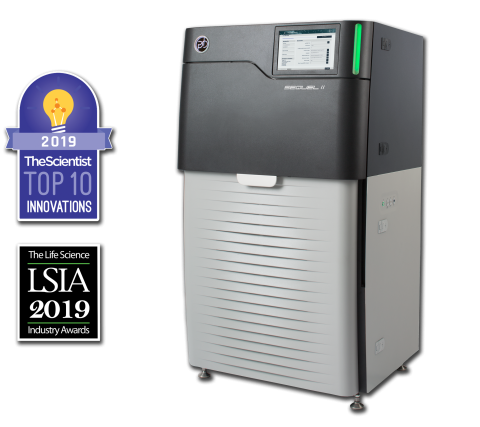
The Sequel II System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by PacBio , and is designed for high throughput, production-scale sequencing laboratories. Originally released in 2015, the Sequel system provides Single Molecule, Real-Time (SMRT) sequencing core...
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCT001 The purpose of this experiment was to evaluate the host responseto West Nile virus (WNV-NY99) wild-type (strain 382) and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain C57BL...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCB001 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain...
Category
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Category
Category
Category
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCN003 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain C57BL...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WDC010 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from primary mouse...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WCN002 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain C57BL...