Filter results
Tags
- (-) Virus (23)
- (-) Soil Microbiology (21)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Viruses (26)
- Omics (25)
- Health (23)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (14)
- Synthetic (14)
- sequencing (13)
- West Nile virus (13)
- Genomics (12)
- Ebola (11)
- Influenza A (11)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Metagenomics (9)
- Resource Metadata (9)
- Microbiome (8)
- Proteomics (8)
- Machine Learning (7)
- Microarray (7)
Systems Virology Lethal Human Virus, SARS-CoV Experiment SM003 New uploads pending The purpose of this SARS experiment was to obtain samples for transcriptome and proteome analysis in C57BlL/6 mice lung infected with dicSARS CoV, SARS MA15 wild type and SARS BatSRBD mutant virus. Overall Design...
Category
Systems Virology Lethal Human Virus, SARS-CoV Experiment SM012 New uploads pending The purpose of this SARS experiment was to obtain samples for transcriptome and proteome analysis in C57Bl/6 mice infected with SARS MA15 or SARS deltaORF6 mutant viruses. Overall Design: Twenty-week-old C57BL/6 mice...
Category
Systems Virology Lethal Human Virus, SARS-CoV Experiment SCL012 New uploads pending The purpose of this SARS experiment was to obtain samples for metabolome and lipidome analysis comparing wild type virus (icSARS) vs icSARS CoV deltaORF6 infected Human lung tissue 2B4 (Calu-3 clonal derivative)...
Category
Systems Virology Lethal Human Virus, SARS-CoV Experiment SHAE002 New uploads pending The purpose of this SARS experiment was to obtain samples for transcriptome analysis in Human Airway Epithelial (HAE) cells infected with SARS-CoV, SARS deltaORF6, SARS BatSRBD mutants in a longitudinal study...
Category
Systems Virology Lethal Human Virus, SARS-CoV Experiment SHAE003 New uploads pending The purpose of this SARS experiment was to obtain samples for transcriptome analysis in Human Airway Epithelial (HAE) cells infected with SARS-CoV, SARS deltaORF6, SARS BatSRBD mutants in a longitudinal study...
Category
Systems Virology Lethal Human Virus, SARS-CoV Experiment SHAE004 New uploads pending The purpose of this SARS experiment was to obtain samples for transcriptome analysis in Human Airway Epithelial (HAE) cells infected with SARS-CoV, SARS deltaORF6, SARS BatSRBD mutants in a longitudinal study...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. Iso-VIG14.1.0 (Metagenome Derived Viral Genomes, WA/IA/KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/IsoVIG14/1770369 Soil samples were collected in triplicate in the Fall of 2017...
Category
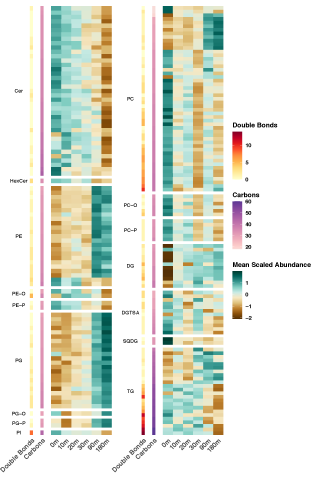
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. IA-TmG.2.0 (Metagenome, IA). [Data Set] PNNL DataHub. https://doi.org/10.25584/IATmG2/1770333 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
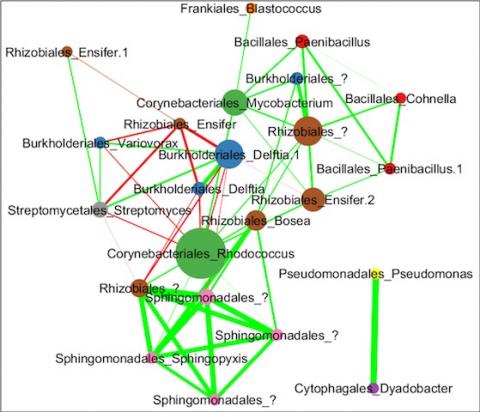
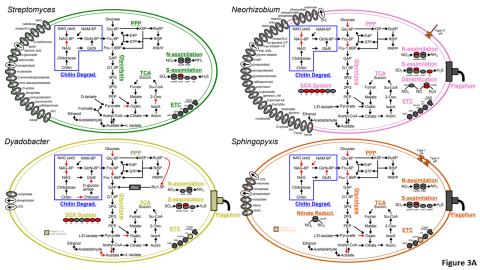
Please cite as : Anderson L.N., J.E. McDermott, and R.S. McClure. 2020. WA-IsoC_MSC1.1.0 (Amplicon 16S rRNA, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WAIsoCMSC1/1635272 The soil microbiome is central to the cycling of carbon and other nutrients and to the promotion of plant growth...
Category
Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. WA-TmG.2.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG2/1770324 Soil samples were collected in triplicate in the fall of 2017 across the three grassland...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. KS-TmG.2.0 (Metagenome, KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/KSTmG2/1770332 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes. [Data Set] PNNL DataHub. https://doi.org/10.25584/PNNLDH/1986536 Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes...