"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Category
- Biology (66)
- Scientific Discovery (66)
- Human Health (52)
- Integrative Omics (48)
- Microbiome Science (11)
- Earth System Science (7)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Plant Science (2)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Materials Science (1)
- National Security (1)
Content type
Tags
- (-) Differential Expression Analysis (46)
- (-) PerCon SFA (10)
- (-) Machine Learning (7)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Viruses (26)
- Omics (25)
- Health (23)
- Virus (23)
- Soil Microbiology (21)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (14)
- Synthetic (14)
- sequencing (13)
- West Nile virus (13)
- Genomics (12)
- Ebola (11)
- Influenza A (11)
- High Throughput Sequencing (9)
- Metagenomics (9)
- Resource Metadata (9)
- Microbiome (8)
- Proteomics (8)
- Microarray (7)
Elmore JR, Dexter GN, Baldino H, Huenemann JD, Francis R, Peabody GL 5th, Martinez-Baird J, Riley LA, Simmons T, Coleman-Derr D, Guss AM, Egbert RG. High-throughput genetic engineering of nonmodel and undomesticated bacteria via iterative site-specific genome integration. Sci Adv. 2023 Mar 10;9(10)...
The recently developed real-time nuclear–electronic orbital (RT-NEO) approach provides an elegant framework for treating electrons and selected nuclei, typically protons, quantum mechanically in nonequilibrium dynamical processes. However, the RT-NEO approach neglects the motion of the other nuclei...
The rhizosphere represents a dynamic and complex interface between plant hosts and the microbial community found in the surrounding soil. While it is recognized that manipulating the rhizosphere has the potential to improve plant fitness and health, engineering the rhizosphere microbiome through...
Agriculture is the largest source of greenhouse gases (GHG) production. Conversion of nitrogen fertilizers into more reduced forms by microbes through a process known as biological nitrification drives GHG production, enhances proliferation of toxic algal blooms, and increases cost of crop...
Metabolite exchange between plant roots and their associated rhizosphere microbiomes underpins plant growth promotion by microbes. Sorghum bicolor is a cereal crop that feeds animals and humans and is used for bioethanol production. Its root tips exude large amounts of a lipophilic benzoquinone...
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
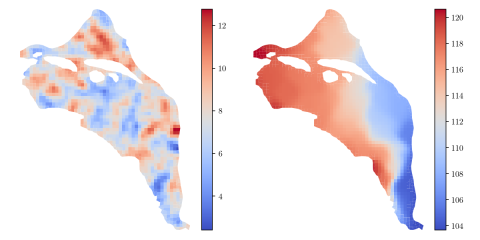
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
Omics Lethal Human Virus, SARS-CoV Experiment SM001 New uploads pending The purpose of this experiment was to evaluate the human host response to Severe Acute Respiratory Syndrome coronavirus (SARS-CoV) wild-type virus. Sample data was obtained for 20 week-old C57BL/6J mouse lung tissue infected...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Ebola Virus Experiment EH001 The purpose of this experiment was to evaluate the human patient peripheral blood mononuclear cells (PBMC) response to Zaire Ebola virus (strain Makona) infection during the 2013-2016 epidemic in West Africa...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL003 The purpose of this experiment was to evaluate the human host response to wild-type infectious clone of Middle Eastern Respiratory Syndrome coronavirus (icMERS-CoV) infection and mockulum. Sample data was obtained...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WSE001 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 virus infection. Sample data was obtained from mouse (strain C57BL6/JAX) blood serum...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WDC010 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from primary mouse...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WGCN002 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WGCN003 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A virus infection. Sample data was obtained from mouse (strain...