This data package consists of data from MIN6 cells (n=4) treated with IL-1β + IFNγ + TNFα for 24 h post-RNAi targeting ADP-ribosylhydrolase ARH3 (Adprhl2), and submitted to proteomics analysis ( Table 1 ). ARH3 is a hydrolase that keeps cellular ADP-ribosylation levels in check and is regulated by...
Filter results
Category
- (-) Integrative Omics (97)
- (-) National Security (32)
- Scientific Discovery (407)
- Biology (288)
- Earth System Science (169)
- Human Health (113)
- Microbiome Science (50)
- Computational Research (25)
- Energy Resiliency (22)
- Computing & Analytics (18)
- Renewable Energy (14)
- Chemical & Biological Signatures Science (12)
- Weapons of Mass Effect (12)
- Materials Science (11)
- Chemistry (10)
- Data Analytics & Machine Learning (9)
- Water Power (8)
- Computational Mathematics & Statistics (7)
- Data Analytics & Machine Learning (7)
- Atmospheric Science (6)
- Ecosystem Science (6)
- Visual Analytics (6)
- Coastal Science (4)
- Energy Storage (4)
- Plant Science (4)
- Solar Energy (4)
- Bioenergy Technologies (3)
- Energy Efficiency (3)
- Transportation (3)
- Cybersecurity (2)
- Distribution (2)
- Electric Grid Modernization (2)
- Grid Cybersecurity (2)
- Subsurface Science (2)
- Wind Energy (2)
- Advanced Lighting (1)
- Computational Mathematics & Statistics (1)
- Environmental Management (1)
- Federal Buildings (1)
- Geothermal Energy (1)
- Grid Analytics (1)
- Grid Energy Storage (1)
- High-Performance Computing (1)
- Terrestrial Aquatics (1)
- Vehicle Technologies (1)
- Waste Processing (1)
Tags
- Virology (55)
- Immune Response (51)
- Time Sampled Measurement Datasets (51)
- Differential Expression Analysis (46)
- Gene expression profile data (45)
- Homo sapiens (38)
- Multi-Omics (33)
- Mass spectrometry data (31)
- Mus musculus (19)
- MERS-CoV (18)
- West Nile virus (13)
- Mass Spectrometry (12)
- Influenza A (11)
- Proteomics (11)
- Ebola (9)
- Omics (9)
- Predictive Phenomics (7)
- Resource Metadata (7)
- Human Interferon (6)
- Microarray (6)
- Omics-LHV Project (6)
- Synthetic (5)
- Mass Spectrometer (4)
- Imaging (3)
- Lethal Human Viruses (3)
- Limited Proteolysis (3)
- metabolomics (3)
- Microscopy (3)
- Response to type I interferon (3)
- TA1 (3)
It consists of human islets (n=10, subdivided into 2 batches) treated with IFNγ + IL-1β for 24 h and submitted to proteomics, lipidomics and metabolomics in the same samples ( Table 1 ) Data contributors: Ernesto S. Nakayasu, Jennifer Kyle, Young-Mo Kim, Soumyadeep Sarkar & Thomas O. Metz...
Category
This data package consists of data from β-TC-6 cells treated with IFNγ + IL-1β for 4, 8 or 24 h (n=4) and submitted to enrichment of cysteine-containing peptides (redox), phosphopeptides and acetylated peptides. This enabled the analysis of proteomics, redox cysteine, phosphoproteomics and...
Category
This data package consists data from EndoC-βH1 (n=6) treated with IFNα for 8 and 24 h and submitted to RNA-seq analysis ( Table 1 ). This was generated in parallel with Pck008. Data contributors: Arturo Roca-Rivada, Xiaoyan Yi & Décio L. Eizirik: ULB Center for Diabetes Research, Medical Faculty...
Category
The data package consists of RNA-seq data from human islet-derived exosomes post IFNγ + IL-1β treatment of human islets (n=5) for 24 h ( Table 1 ). Data contributors: Preethi Krishnan& Carmella Evans-Molina: Department of Diabetes Immunology, Beckman Research Institute, City of Hope, Duarte, CA, USA...
Category
This data package consists of data from paired human islets treated with IFNγ + IL-1β for 48 h and submitted to RNA-seq (n=5), ATAC-seq (n=5), UMI-4C18 (n=2 or 3, depending on the viewpoint), and ChIP-seq for histone H3 lysine 27 acetylation (H3K27Ac) (n=4) ( Table 1 ). Islets across the study...
Category
This data package consists of data from MIN6 cells treated with IL-1β + IFNγ + TNFα + palmitate for 24 h and submitted to proteomics analysis, and the same treatment for 8 h for metabolomics analysis (n=4) ( Table 1 ). This experimental design was set to study the possible roles of free fatty acids...
Category
This data package contains processed LC-MS proteomics results and analysis scripts associated with the paper "Assessing Degenerate Peptide Resolution Methods using a Ground Truth Dataset". In this study, we designed an artificial microbial community to create a ground truth, simulated metaproteomic...
Category
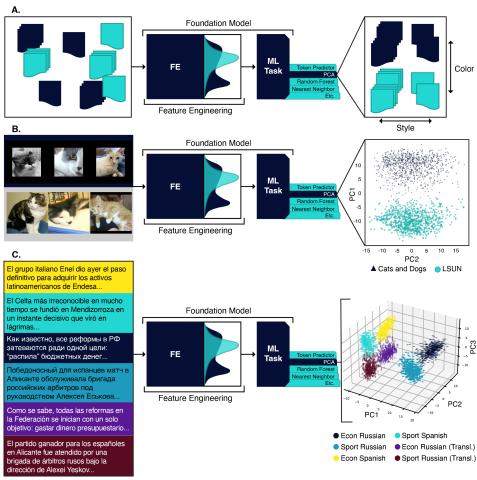
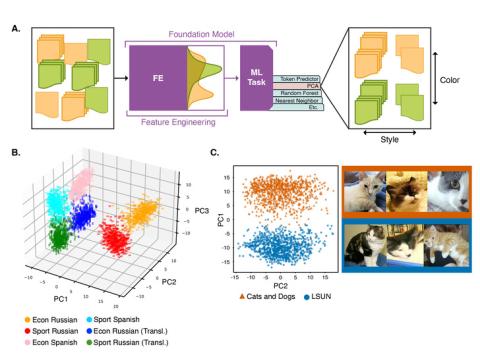
This repository contains data and code for the experiments run in the paper "Understanding Generative AI Content with Embedding Models" ( https://arxiv.org/abs/2408.10437 ). DataBase POC: Max Vargas (max.vargas@pnnl.gov) The data is separated by experiment: A. The `stack_exchange` dataset contains a...
Category
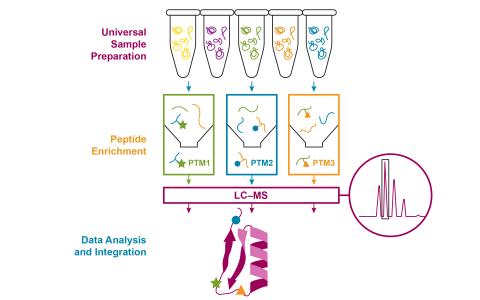
Dataset Description The goal of the experiment was to demonstrate that the optimized multiplexed multi-PTM profiling workflow can comprehensively and quantitatively capture dynamic changes in protein abundance, cysteine oxidation, phosphorylation, and acetylation in cytokine-induced inflammatory...
Category
This repository contains data for the experiments run in the paper "Understanding Generative AI Content with Embedding Models" ( https://arxiv.org/abs/2408.10437 ). DataBase POC: Max Vargas (max.vargas@pnnl.gov) The data is separated by experiment: A. The `stack_exchange` dataset contains a...
Category
Currently pending public release.
Currently pending public release.
Created on 2024-10-16T17:27:40+00:00 by LN Anderson ; Last updated 2025-08-01T15:13:46+00:00. S. elongatus PCC 7942 LiP and TPP Structural Proteomics (JM-PB-DP3) The purpose of this experiment was to investigate structural alterations in proteins involved in central carbon metabolism and...
Dataset Description The purpose of this experiment was to evaluate the human host cellular response to wild type human coronavirus strain 229E (HCoV-229E) infection. Sample data was obtained for mock-infected and immortalized human lung fibroblasts (MRC-5) (MOI 5). Whole cell lysates were collected...